Characterization of kinase gene expression and splicing profile in prostate cancer with RNA-Seq data
Abstract
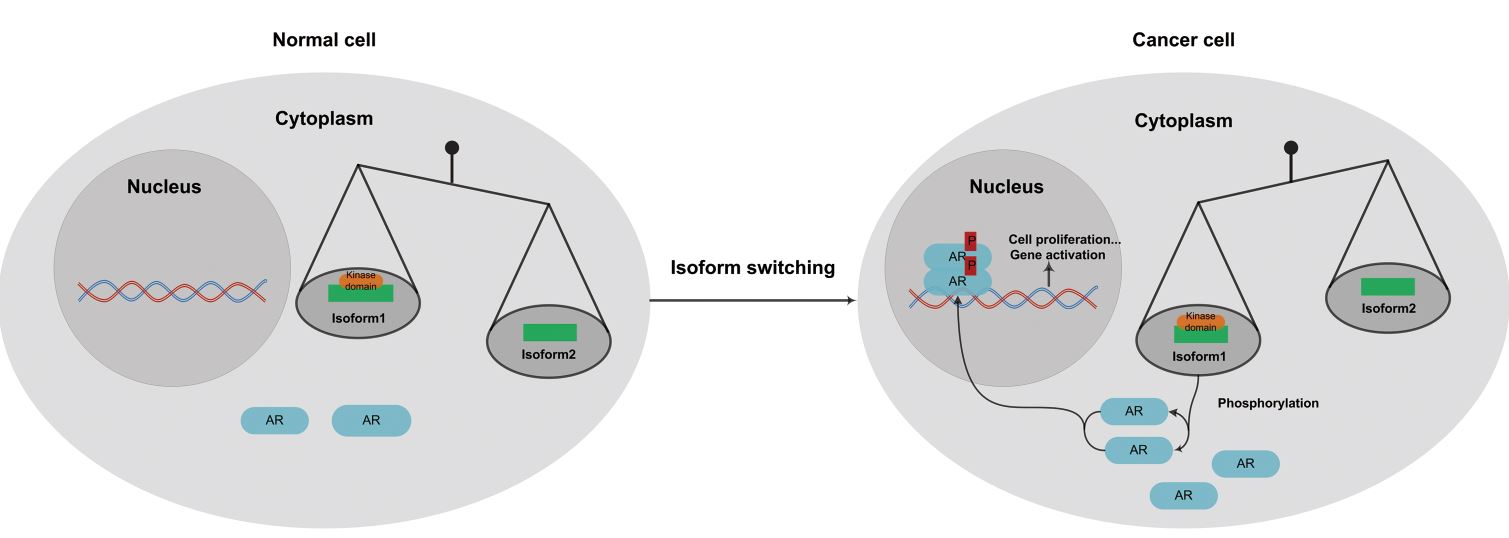
Alternative splicing is a ubiquitous post-transcriptional process in most eukaryotic genes. Aberrant splicing isoforms and abnormal isoform ratios can contribute to cancer development. Kinase genes are key regulators of various cellular processes. Many kinases are found to be oncogenic and have been intensively investigated in the study of cancer and drugs. RNA-Seq provides a powerful technology for genome-wide study of alternative splicing in cancer besides the conventional gene expression profiling. But this potential has not been fully demonstrated yet. Here we characterized the transcriptome profile of prostate cancer using RNA-Seq data from viewpoints of both differential expression and differential splicing, with an emphasis on kinase genes and their splicing variations. We built up a pipeline to conduct differential expression and differential splicing analysis. Further functional enrichment analysis was performed to explore functional interpretation of the genes. With focus on kinase genes, we performed kinase domain analysis to identify the functionally important candidate kinase gene in prostate cancer. We further calculated the expression level of isoforms to explore the function of isoform switching of kinase genes in prostate cancer. We identified distinct gene groups from differential expression and splicing analysis, which suggested that alternative splicing adds another level to gene expression regulation. Enriched GO terms of differentially expressed and spliced kinase genes were found to play different roles in regulation of cellular metabolism. Function analysis on differentially spliced kinase genes showed that differentially spliced exons of these genes are significantly enriched in protein kinase domains. Among them, we found that gene CDK5 has isoform switching between prostate cancer and benign tissues, which may affect cancer development by changing androgen receptor (AR) phosphorylation. The observation was validated in another RNA-Seq dataset of prostate cancer cell lines. Our work characterized the expression and splicing profile of kinase genes in prostate cancer and proposed a hypothetical model on isoform switching of CDK5 and AR phosphorylation in prostate cancer. These findings bring new understanding to the role of alternatively spliced kinases in prostate cancer and demonstrate the use of RNA-Seq data in studying alternative splicing in cancer.