Modeling RNA-binding protein specificity in vivo by precisely registering protein-RNA crosslink sites
Abstract
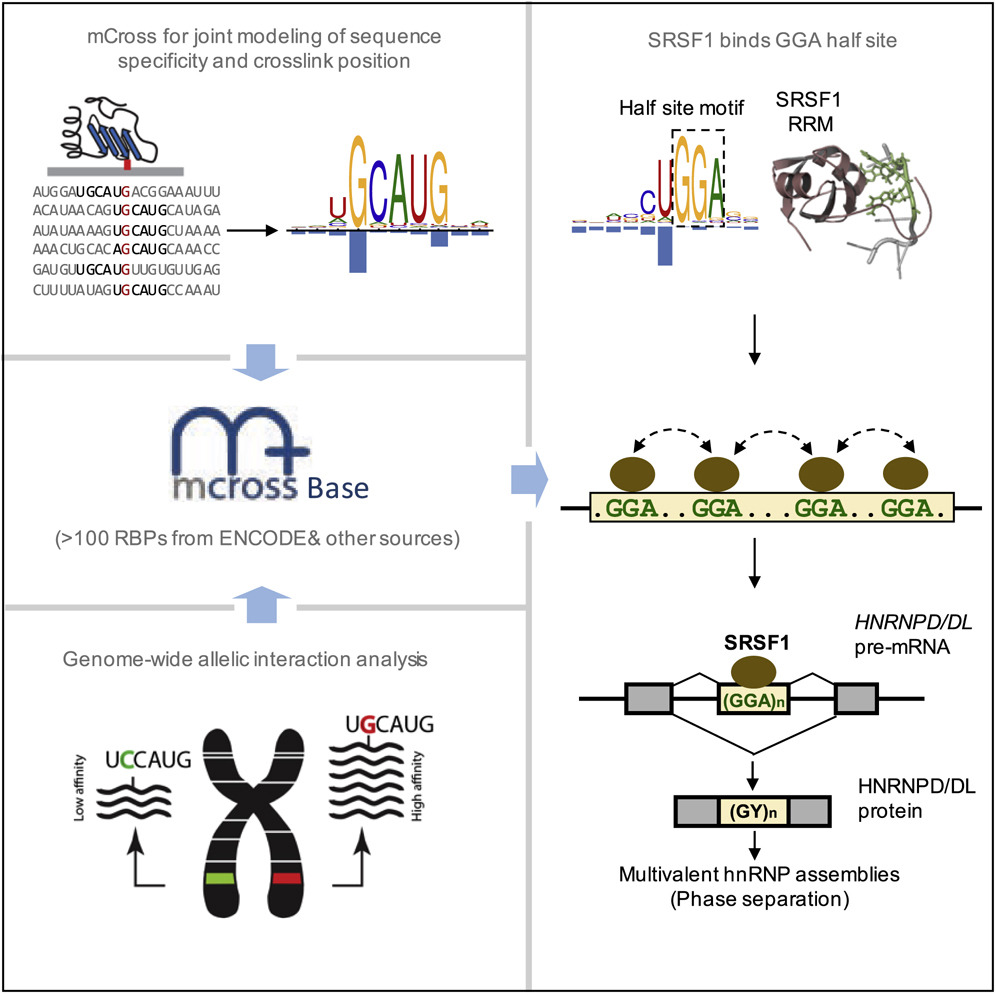
RNA-binding proteins (RBPs) regulate post-transcriptional gene expression by recognizing short and degenerate sequence motifs in their target transcripts, but precisely defining their binding specificity remains challenging. Crosslinking and immunoprecipitation (CLIP) allows for mapping of the exact protein-RNA crosslink sites, which frequently reside at specific positions in RBP motifs at single-nucleotide resolution. Here, we have developed a computational method, named mCross, to jointly model RBP binding specificity while precisely registering the crosslinking position in motif sites. We applied mCross to 112 RBPs using ENCODE eCLIP data and validated the reliability of the discovered motifs by genome-wide analysis of allelic binding sites. Our analyses revealed that the prototypical SR protein SRSF1 recognizes clusters of GGA half-sites in addition to its canonical GGAGGA motif. Therefore, SRSF1 regulates splicing of a much larger repertoire of transcripts than previously appreciated, including HNRNPD and HNRNPDL, which are involved in multivalent protein assemblies and phase separation.